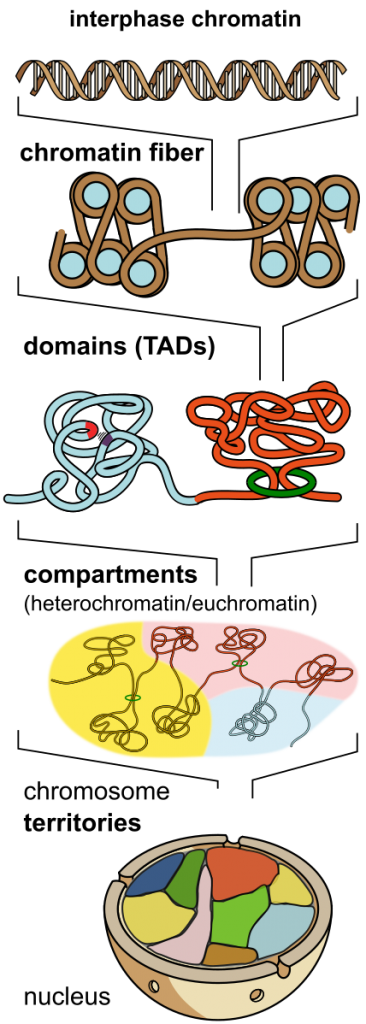
Chromatin represents the physical form of the linear genomic information that provides the blueprint for every eukaryotic cell. While the extremely long and fine strings of DNA store the linear information of genes and controlling elements, the protein component of chromatin provides protection and functional organization. Current and past research efforts are focused with great energy on conceptualizing genome architecture (Figure 1) and on relating the physical structure of the genome to function. At the primary structural level genomes are stored in the form of nucleosome arrays, the chromatin fibre, which control access to the underlying DNA sequence. Despite decades of research, the impact of chromatin fibre architecture produced by nucleosome interactions is poorly understood, particularly in vivo. While chromatin fibres form regular structures in isolation, these structures do not seem to be prevalent in regular interphase cells. However, several lines of evidence argue that chromatin higher-order structure is not completely random.
We are interested in understanding the molecular interactions that govern chromatin structure at the highest possible resolution. We, therefore, study chromatin arrays and are trying to understand how nucleosomes interact with each other and with regulatory factors.